The code that goes with this post is at https://github.com/Atrix256/OT1D.
The title of this post is a mouthful but the topic is pretty useful.
Let’s say that we have two histograms:
- 1,2,3,1,0,0,0,0,0,0
- 0,0,0,0,0,0,1,2,3,1
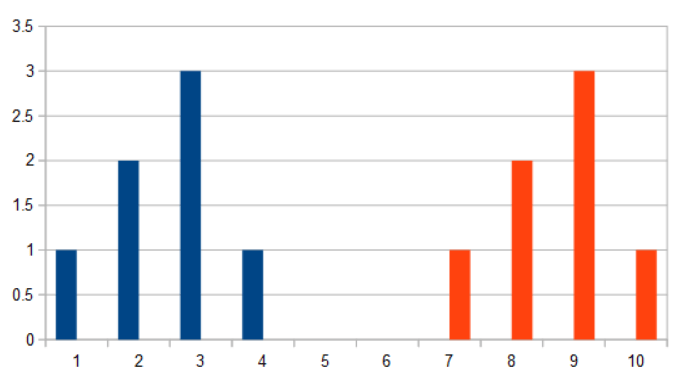
When you want to see how similar two histograms are, some common ways are:
- Treat the histograms as high dimensional vectors and dot product them. Higher values mean more similar.
- Treat the histograms as high dimensional points and calculate the distance between them, using the regular distance formula (aka L2 norm or Euclidian norm), taxi cab distance (aka L1 norm) or others. Lower values mean more similar.
Those aren’t bad methods, but they aren’t always the right choice depending on your needs.
- If you dot product these histograms as 10 dimensional vectors, you get a value of zero because there’s no overlap. If there are more or fewer zeros between them, you still get zero. This gives you no idea how similar they are other than they don’t overlap.
- If you calculate the distance between these 10 dimensional points, you will get a non zero value which is nice, but adding or removing zeros between them, the value doesn’t change. You can see this how the length of the vector (1,0,1) is the square root of 2, which is also the length of the vector (1,0,0,1) and (1,0,0,0,1). It doesn’t tell you really how close they are on the x axis.
In this case, let’s say we want a similarity metric where if we separate these histograms with more zeros, it tells us that they are less similar, and if we remove zeros or make them overlap, it tells us that they are more similar.
This is what the p-Wasserstein distance gives us. When we interpolate along this distance metric from the first histogram to the second, the first histogram slides to the right, and this is actually 1D optimal transport. This is in contrast to us just linearly interpolating the values of these histograms, where the first histogram would shrink as the second histogram grew.
The title of this post says that we will be working with both histograms and PDFs (probability distribution functions). A PDF is sort of like a continuous version of a histogram, but integrates to 1 and is limited to non negative values. In fact, if a histogram has only non negative values, and sums up to 1, it can be regarded as a PMF, or probability mass function, which is the discrete version of a PDF and is used for things like dice rolls, coin flips, and other discrete valued random events.
You can do similar operations with PDFs as you can with histograms to calculate similarity.
- Instead of calculating a dot product between vectors, you take the integral of the product of the two functions. Higher values means more similar, like with dot product.
. This is a a kind of continuous dot product.
- Instead of calculating a distance between points, you take the integral of the absolute value of the difference of the two functions raised to some power and then the result taken to one over the same power
. Lower values means more similar, like with distance. This is a kind of continuous distance calculation.
Also, any function that doesn’t have negative y values can be normalized to integrate to 1 over a range of x values and become a PDF. So, when we talk about PDFs in this post, we are really talking about any function that doesn’t have negative values over a specific range.
p-Wasserstein Distance Formula
“Optimal Transport: A Crash Course” by Kolouri et. al (https://www.imagedatascience.com/transport/OTCrashCourse.pdf) page 46 says that you can calculate p-Wasserstein distance like this (they use different symbols):
That looks a bit like the continuous distance integral we saw in the last section, but there are some important differences.
Let me explain the symbols in this function. First is which is where the p in p-Wasserstein distance comes from. It can be 1, 2, 1.5, or any other number. If using the value 2, you’d call it 2-Wasserstein distance, and it would be dealing with the squares of differences between these functions
and
.
Let’s say that we have a PDF called f. We can integrate that to get a CDF or cumulative distribution function that we’ll call F. We can then switch x and y and solve for y to get the inverse of that function, or . If we have another PDF called g, we can follow the same process to get
.
When we calculate the integral above, either analytically / symbolically if we are able, or numerically otherwise, we get the p-Wasserstein distance which has the nice properties we described before.
A fun tangent: if you have the inverse CDF of a PDF, you can plug in a uniform random number in [0,1] to the inverse CDF and get a random number drawn from the original PDF. It isn’t always possible or easy to get the inverse CDF of a PDF, but this also works numerically – you can make a table out of the PDF, make a table for the CDF, and make a table for the inverse CDF from that, and do the same thing. More info on all of that at https://blog.demofox.org/2017/08/05/generating-random-numbers-from-a-specific-distribution-by-inverting-the-cdf/.
Calculating 2-Wasserstein Distance of Simple PDFs
Let’s start off by calculating some inverse CDFs of simple PDFs, then we’ll use them in the formula for calculating 2-Wasserstein distances.
Uniform PDF
First is the PDF , which is for uniform random values. It integrates to 1 for x between 0 and 1 and is positive in that range as well.
If we integrate that to get the CDF, we get .
If we flip x and y, and solve again for y to get the inverse CDF, we get .
Linear PDF
The next PDF is . It integrates to 1 for x between 0 and 1 and is positive in that range as well.
If we integrate that, the CDF is .
Flipping x and y and solving for y gives us the inverse CDF .
Quadratic PDF
The last PDF is . It integrates to 1 for x between 0 and 1 and is positive in that range as well.
If we integrate that, the CDF is
Flipping x and y and solving for y gives us the inverse CDF .
2-Wasserstein Distance From Uniform to Linear
The 2-Wasserstein distance formula looks like this:
Let’s square the function for now and we’ll square root the answer later. Let’s also replace with the uniform PDF inverse CDF, and replace
with the linear PDF inverse CDF. We can drop the absolute value since the square does that for us.
Expanding the equation, we get:
Luckily we can break that integral up into 3 integrals:
https://www.wolframalpha.com/ comes to the rescue here where you can ask it “integrate y=x^2 from 0 to 1” etc to get the answer to each integral.
Remembering that we removed the square root from the formula at the beginning, we have to square root this to get our real answer.
2-Wasserstein Distance From Uniform to Quadratic
We can do this again measuring the distance between uniform and quadratic.
Square rooting that gives:
2-Wasserstein Distance From Linear to Quadratic
We can do this again between linear and quadratic.
Square rooting that gives:
These results tell us that uniform and quadratic are most different, and that linear and quadratic are most similar. Uniform and linear are in the middle. That matches what you’d expect when looking at the graphs.

Calculating p-Wasserstein Distance of More Complicated PDFs
If you have a more complex PDF that you can’t make an inverted CDF of, you can still calculate the distance metric numerically.
You start off by making a table of PDF values at evenly spaced intervals, then divide all entries by the sum of all entries to make it a tabular form of the PDF (it sums to 1). You then make a CDF table where each entry is the sum of PDF values at or before that location.
Now to evaluate the inverted CDF for a value x, you can do a binary search of the CDF table to find where that value x is. You also calculate what percentage the value is between the two indices it’s between so that you have a better approximation of the inverted CDF value. Alternately to a binary search, you could make an ICDF table instead.
If that sounds confusing, just imagine you have an array where the x axis is the array index and the y axis is the value in the array. This is like a function where if you have some index you can plug in as an x value, it gives you a y axis result. That is what the CDF table is. If you want to evaluate the inverted CDF or ICDF, you have a y value, and you are trying to find what x index it is at. Since the y value may be between two index values, you can give a fractional index answer to make the ICDF calculation more accurate.
Now that you can make an inverted CDF table of any PDF, we can use Monte Carlo integration to calculate our distance metric.
Let’s look at the formula again:
To calculate the inner integral, we pick a bunch of x values between 0 and 1, plug them into the equation and average the results that come out. After that, we take the answer to the 1/p power to get our final answer. The more samples you take of x, the more accurate the answer is.
Here is some C++ that implements that, with the ICDF functions being template parameters so you can pass any callable thing in (lambda, function pointer, etc) for evaluating the ICDF.
template <typename ICDF1, typename ICDF2>
float PWassersteinDistance(float p, const ICDF1& icdf1fn, const ICDF2& icdf2fn, int numSamples = 10000000)
{
// https://www.imagedatascience.com/transport/OTCrashCourse.pdf page 45
// Integral from 0 to 1 of abs(ICDF1(x) - ICDF2(x))^p
// Then take the result to ^(1/p)
double ret = 0.0;
for (int i = 0; i < numSamples; ++i)
{
float x = RandomFloat01();
float icdf1 = icdf1fn(x);
float icdf2 = icdf2fn(x);
double y = std::pow(std::abs((double)icdf1 - (double)icdf2), p);
ret = Lerp(ret, y, 1.0 / double(i + 1));
}
return (float)std::pow(ret, 1.0f / p);
}
If both ICDFs are tables, that means they are both piecewise linear functions. You can get a noise free, deterministic calculation of the integral by integrating the individual sections of the tables, but the above is a generic way that will work for any (and mixed) methods for evaluating ICDFs.
Below is a table of the 2-Wasserstein distances calculated different ways. The analytic truth column is what we calculated in the last section and is the correct answer. The numeric function column is us running the above numerical integration code, using the inverted CDF we made symbolically in the last section. The last column “numeric table” is where we made a tabular inverted CDF by taking 10,000 PDF samples and making a 100 entry CDF, then using binary search to do inverse CDF evaluations.
| Analytic Truth | Numeric Function | Numeric Table | |
| Uniform To Linear | 0.18257418583 | 0.182568 | 0.182596 |
| Uniform To Quadratic | 0.27602622373 | 0.276012 | 0.275963 |
| Linear To Quadratic | 0.09534625892 | 0.095327 | 0.095353 |
For kicks, here are p values of 1 and 3 to compare against 2. Calculated using the “numeric table” method.
| 1 | 2 | 3 | |
| Uniform To Linear | 0.166609 | 0.182596 | 0.192582 |
| Uniform To Quadratic | 0.249933 | 0.275963 | 0.292368 |
| Linear To Quadratic | 0.083308 | 0.095353 | 0.103187 |
Calculating p-Wasserstein Distance of PMFs
If you have a random variable with discrete random values – like a coin toss, or the roll of a 6 sided dice – you start with a PMF (probability mass function) instead of a PDF. A PMF says how likely each discrete event is, and all the events add up to 1.0. The PMF is probably already a table. You can then turn that into a CDF and continue on like we did in the last section. Let’s see how to make a CDF for a PMF.
When you evaluate y=CDF(x), the y value you get is the probability of getting the value x or lower. When you turn a PMF into a CDF, since it’s discrete, you end up with a piecewise function.
For instance, here is the the PMF 0.1, 0.3, 0.2, 0.4.
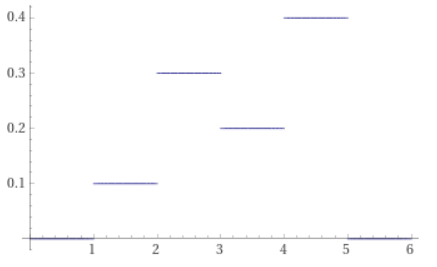
The CDF is 0.1, 0.4, 0.6, 1.0.
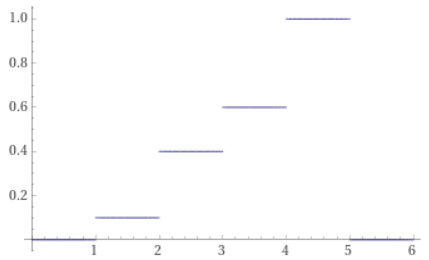
With a CDF you can now do the same steps as described in the last section.
Interpolating PDFs & Other Functions
Interpolating over p-Wassertstein distance is challenging in the general case. I’m going to dodge that for now, and just show the nice and easy way to interpolate over 1-Wasserstein distance.
First let’s see what happens if we interpolate two gaussian PDFs directly… by linearly interpolating PDF(x) for each x to make a new PDF. As you can see, the gaussian on the left shrinks as the gaussian on the right grows.
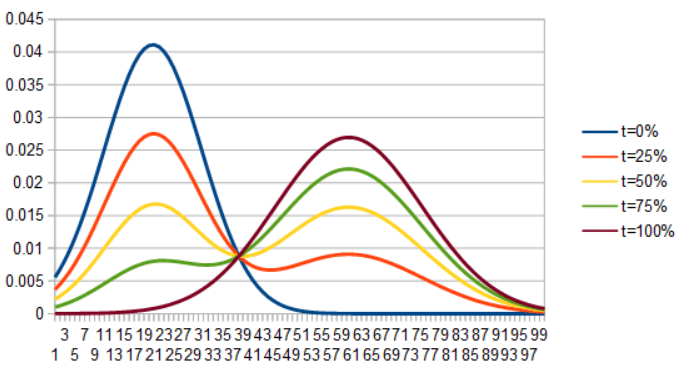
If we instead lerp ICDF(x) for each x to make a new ICDF, invert it to make a CDF, then differentiate it to get a PDF, we get this very interesting optimal transport interpolation:
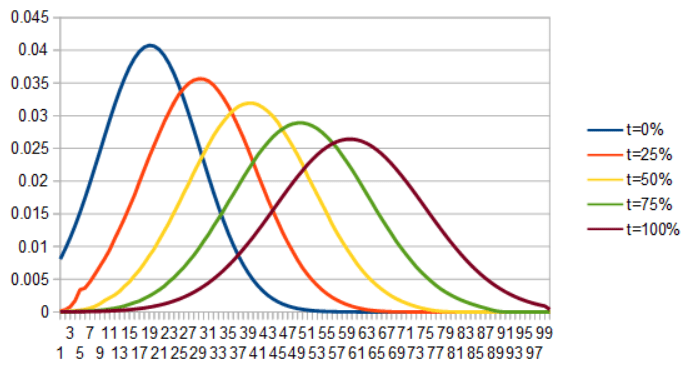
For a second example here is a uniform PDF interpolating into Gaussian, by lerping the PDFs and then the ICDFs.
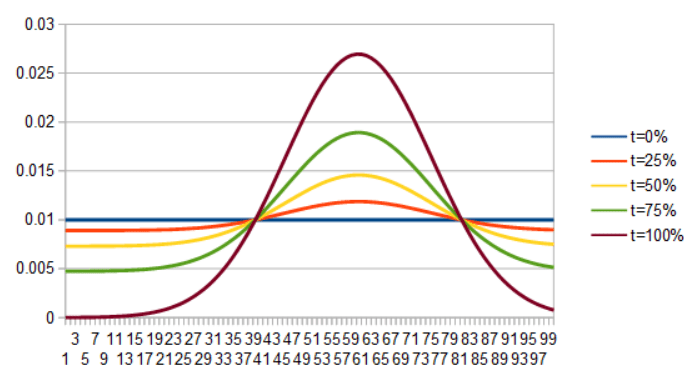
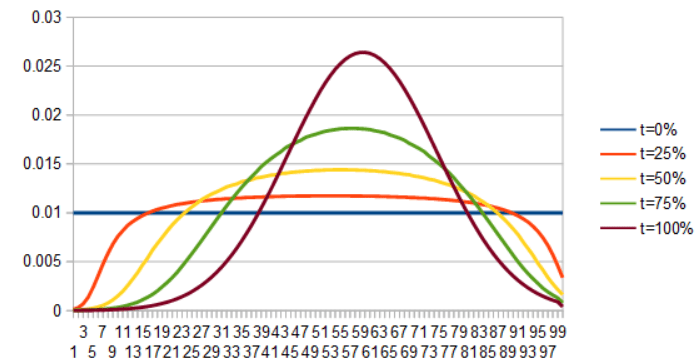
As a last set of examples, here is uniform to quadratic. First by lerping the PDFs, then by lerping the ICDFs.

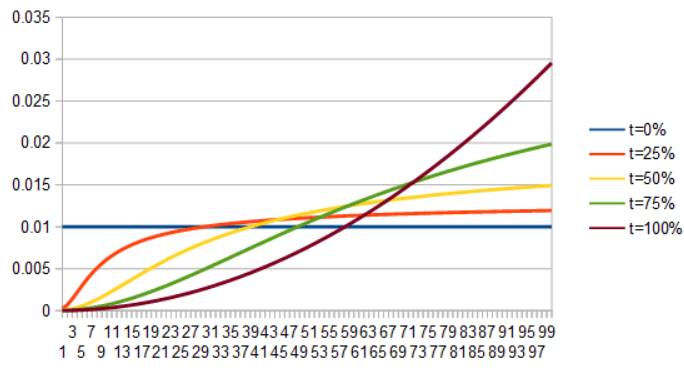
It sounds like a lot of work interpolating functions by integrating the PDFs or PMFs to CDFs, inverting them to ICDFs, lerping, then inverting and differentiating, and it really is a lot of work.
However, like I mentioned before, if you a plug a uniform random value into an ICDF function, it will give you an unbiased random value from the PDF. This means that in practice, if you are generating non uniform random numbers, you often have an ICDF handy for the distribution already; either symbolically or numerically in a table. That makes this whole process a lot simpler: plug your uniform random x value into the two ICDFs, and return the result of lerping them. That’s all there is to it to generate random numbers that are from a PDF interpolated over 1-Wasserstein distance. That is extremely simple and very runtime friendly for graphics and game dev people. Again, more simply: just generate 2 random numbers and interpolate them.
If you are looking for some intuition as to why interpolating the ICDF does what it does, here are some bullet points.
- PDF(x) or PMF(x) tells you how probable x is.
- CDF(x) tells you how probable it is to get the value x or lower.
- ICDF(x) tells you the value you should get, for a uniform random number x.
- Interpolating PDF1(x) to PDF2(x) makes the values in PDF1 less likely and the values in PDF2 more likely. It’s shrinking PDF1 on the graph, and growing PDF2.
- Interpolating CDF1(x) to CDF2(x) gets the values that the two PDFs would give you for probability x, and interpolates between them. If a random roll would get you a 3 from the first PDF, and a 5 from the second PDF, this interpolates between the 3 and the 5.
In this section we’ve talked about linear interpolation between two PDFs, but this can actually be generalized.
- A linear interpolation is just a degree 1 (linear) Bezier curve, which has 2 control points that are interpolated between. The formula for a linear Bezier curve is just A(1-t)+Bt which looks exactly like linear interpolation, because it is! With a quadratic Bezier curve, you have 3 control points, which means you could have three probability distributions you were interpolating between, but it would only pass through the two end points – the middle PDF would never be fully used, it would just influence the interpolation. You can do cubic and higher as well for more PDFs to interpolate between. Check this out for more info about Bezier curves. https://blog.demofox.org/2015/05/25/easy-binomial-expansion-bezier-curve-formulas/.
- A linear interpolation is a Bezier curve on a one dimensional 1-simplex aka a line with barycentric coordinates of (s,t) where s=(1-t). The formula for turning barycentric coordinates into a value is As+Bt which is again just linear interpolation. You can take this to higher dimensions though. In two dimensions, you have a 2-simple aka a triangle with barycentric coordinates of (u,v,w). The formula for getting a value on a (linear) Bezier triangle is Au+Bv+Cw where A, B and C would be 3 different PDFs in this case. Higher order Bezier triangles exist too. Have a look here for more information: https://blog.demofox.org/2019/12/07/bezier-triangles/
- Lastly, Bezier surfaces and (hyper) volumes exist, if you want to interpolate in rectangular domains aka “tensor product surfaces”. More info on that here: https://blog.demofox.org/2014/03/22/bezier-curves-part-2-and-bezier-surfaces/.
To see how to linearly interpolate in other p-Wasserstein distance metrics, give this link below a read. I’ll be doing a post on optimal transport in higher dimensions before too long and will revisit it then.
https://pythonot.github.io/auto_examples/plot_barycenter_1D.html
Thanks for reading!